 |
|
|
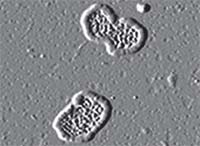
The disease-causing variant of myocilin forms large circular fibrils.
|
This is the first time these molecules have been identified; the discovery signifies potential drug targets for glaucoma, according to Georgia Institute of Technology researchers.
In the collaborative study with Emory University and the University of South Florida, the Georgia Tech researchers designed a simple, high-throughput assay and screened a library of compounds.1 They identified two molecules that bound tightly to the mutant protein myocilin, which aggregates within trabecular meshwork cells.
The researchers then added these molecules to cultured human cells that make the toxic aggregating myocilin. They found that treating the cells with the newly identified molecules not only blocked the mutant myocilin from aggregating, but also released mutant myocilin from the cells, reducing toxicity.
“When we saw that these compounds inhibited aggregation, then we knew we were onto something good because aggregation underlies the pathogenesis of this form of glaucoma,” says principal investigator Raquel Lieberman, PhD.
In a separate study, the researchers found that myocilin aggregates are similar to amyloid protein deposits, which are responsible for Alzheimer’s disease and other neurodegenerative diseases.2 “In Alzheimer’s disease, the deposits are extracellular and kill neurons. In glaucoma, the aggregates are not directly killing neurons in the retina to cause vision loss, but they are cytotoxic in the pressure-regulating region of the eye,” Dr. Lieberman says. “It’s parallel to all these other amyloids that are out there in neurodegenerative disease.” The researchers are now learning more about what myocilin does and why it’s in the eye in the first place.
1. Orwig SD, Chi PV, Du Y, et al. Ligands for glaucoma-associated myocilin discovered by a generic binding assay. ACS Chem Biol. 2013 Dec 9. [Epub ahead of print]
2. Hill SE, Donegan RK, Lieberman RL. The glaucoma-associated olfactomedin domain of myocilin forms polymorphic fibrils that are constrained by partial unfolding and Peptide sequence. J Mol Biol. 2014 Feb 20;426(4):921-35.

